What We Do
Aethron Labs is an independent research lab focused on developing large-scale machine learning systems for interpreting complex scientific data. Our work is centered on building foundational capabilities rather than narrow tools or application-specific models.
Progress Log
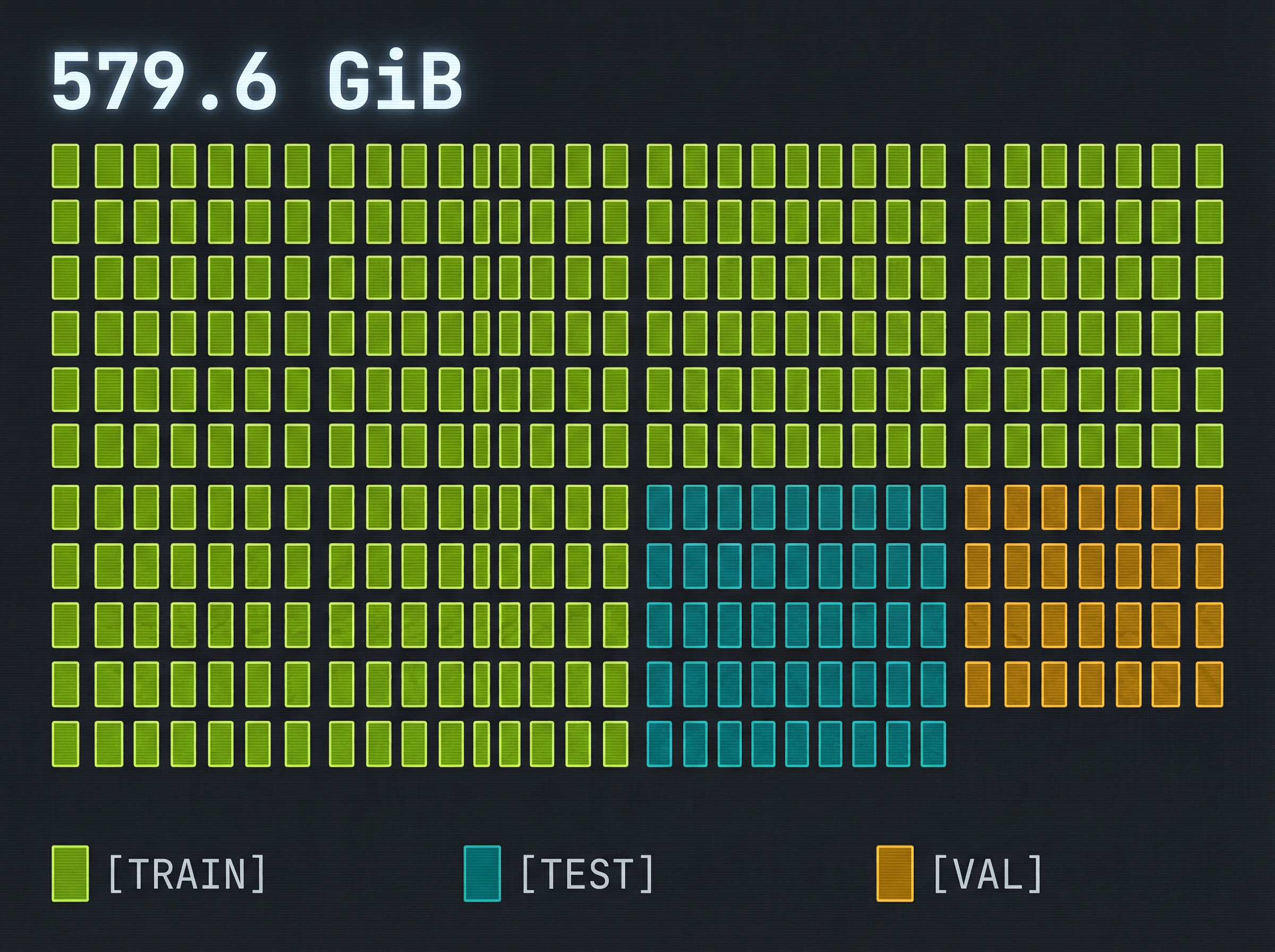
- ◈Acquired large-scale MS/MS dataset (~579 GiB)
- ◈Built Rust + Python preprocessing pipeline
- ◈Achieved ~160K spectra/sec on commodity CPUs
- ◈Processed 100% into verified, versioned shards (GeMS v1)
- ◈Strict train/test/validation splits — no leakage
- ◈Arrow-based training shards + HDF5 workflows
- ◈Trained Final_V3 encoder-only transformer (12.38M params)
- ◈Phase-1 campaign: V1–V3 over ~201M spectra
- ◈Contrastive loss: 0.6206 → 0.3373 (45.7% reduction)
- ◈Validation loss fell 74.2% across V1–V3
- ◈Embedding std stable at 0.072 — promoted Final_V3
- ◈Hardware: 2× RTX PRO 6000 Blackwell, W&B observability
- ◈Identified CROs as primary commercial segment
- ◈Drafted GTM messaging and LOI templates
- ◈Conducted direct CRO outreach
- ◈Applied to BoostVC, Convergent, Artizen, Founders Inc
- ◈Written technical docs and 1-2 page proposals
- ◈Capital-efficient ignition → pre-seed execution plan
GeMS v1 — Dataset Complete
- ◈Rust + Python hybrid preprocessing
- ◈Arrow-format training shards
- ◈Strict no-leakage train/test/val splits
- ◈Versioned, reproducible shard generation
- ◈~18K spectra/sec processing throughput
- ◈Instrument-agnostic normalization
- ◈HuggingFace → VM → storage pipeline
- ◈Verified checksums on all 338 shards
Final_V3 — Foundation Encoder
- ◈Encoder-only transformer, 12.38M parameters
- ◈Self-supervised contrastive pretraining
- ◈Phase-gated training with promotion gates
- ◈W&B observability across all runs
- ◈2× RTX PRO 6000 Blackwell training lane
- ◈Object-storage-first artifacts (Wasabi)
- ◈Resumable hydration + role-specific GPU containers
- ◈Full checkpoint lineage and manifest discipline
V26 — Structure Alignment
- ◈Encoder attaches to RDKit Morgan targets without collapse
- ◈Stable embedding variance across alignment run
- ◈20-example validation gallery: 11/20 ground-truth matches
- ◈Fingerprint decoding shows usable structure signal
- ◈Trained retrieval projection remains weak
- ◈Top-1 ranking not yet reliable
- ◈Confidence calibration negative — not promoted
- ◈Next: reranking, ambiguity tiers, confidence-as-abstention
Inference Layer — Qdrant Atlas
- ◈Qdrant vector indexing — nearest-neighbor retrieval
- ◈1M / 5M density atlases for chemical-space inspection
- ◈Outlier triage and rare-compound detection
- ◈Family-analysis surfaces for compound clustering
- ◈4,107 InChIKey prefixes in rich atlas
- ◈Candidate narrowing, not autonomous identification
- ◈Structure-elucidation support for analyst workflows
- ◈Evidence aggregation over ambiguous candidate sets
- ◈Not a de novo generator — bounded, defensible claims
- ◈Ranking and confidence remain the next engineering target
The Problem
Across the life sciences and molecular research, data generation has dramatically outpaced our ability to interpret it. Core analytical technologies produce enormous volumes of rich, high-dimensional measurements, yet downstream understanding still depends on fragile heuristics, limited reference data, and manual analysis.
This gap constrains discovery, slows research, and limits what can be reliably inferred from experimental data.
Our Approach
We believe this is fundamentally a representation problem. Aethron Labs is building foundation models that learn directly from raw scientific data, capturing underlying structure in a way that generalizes across instruments, conditions, and experimental settings.
The goal is not to replace existing workflows, but to create a new computational substrate that makes scientific interpretation more scalable, reliable, and extensible.
Market Opportunity
Mid-to-large CROs typically operate 10s–100s of LC-MS/MS instruments processing millions of spectra per year, with teams of analysts whose time is the primary cost driver. This spend is recurring, operational, and directly tied to throughput and turnaround time.
Initial commercialization targets enterprise API licensing priced against analyst time and throughput. Targeting ~200–500 CROs and pharma analytical groups globally, with early adopters likely the top 10–50 CROs by analytical volume. Initial contracts plausibly in the $100K–$1M ARR range per customer — supporting a credible $50–200M serviceable obtainable market before broader expansion.
Go-To-Market
Simple and Credible
The initial GTM is intentionally narrow and execution-driven. Aethron Labs targets CROs first — the organizations that feel the MS/MS bottleneck most acutely.
Turnaround time, analyst throughput, and defensibility of results directly determine their margins and competitiveness. The goal is not rapid scaling at first, but credible proof that this infrastructure works in real workflows.
What This Becomes
What begins as programmatic molecular search for LC-MS/MS expands as models and representations mature:
- ◈LC-MS/MS annotation and search
- ◈Metabolomics and impurity identification
- ◈Direct integration into CRO and pharma pipelines
- ◈Drug discovery, DMPK, metabolomics, materials research
- ◈Standard interpretation layer — not a standalone tool
- ◈Cross-instrument generalization
- ◈Reusable substrate for molecular and materials science
- ◈TAM expands to multiple tens of billions
- ◈Core scientific computing infrastructure
Founder Profile
- ◈Molecular science
- ◈Biomaterials
- ◈Quantum systems
- ◈Computational fluid dynamics
- ◈Startups
- ◈Large-scale production systems
- ◈Infrastructure engineering
This background spans the full stack required for this problem: scientific domain understanding, large-scale ML systems, and production engineering realities. Aethron Labs is structured to reflect this combination from day one.
Motivation
This effort is motivated by a rare convergence:
Scientific fields are generating orders of magnitude more data
Interpretation remains the bottleneck — not collection
Modern ML can now operate at the scale and complexity required
The opportunity is not incremental optimization. It is to define a new category of scientific infrastructure that sits between raw experimental data and downstream discovery.
By starting with a concrete, economically grounded use case (CRO workflows) and expanding deliberately, Aethron Labs aims to accelerate scientific discovery, improve reproducibility, and create durable infrastructure with impact beyond a single domain.
This is a long-term bet on advancing science as a system, not just improving a workflow.